SelfPacedEnsembleClassifier
- class imbens.ensemble.SelfPacedEnsembleClassifier(estimator=None, n_estimators: int = 50, *, k_bins: int = 5, soft_resample_flag: bool = False, replacement: bool = True, estimator_params=(), n_jobs=None, random_state=None, verbose=0)
A self-paced ensemble (SPE) Classifier for class-imbalanced learning.
Self-paced Ensemble (SPE) [1] is an ensemble learning framework for massive highly imbalanced classification. It is an easy-to-use solution to class-imbalanced problems, features outstanding computing efficiency, good performance, and wide compatibility with different learning models.
This implementation extends SPE to support multi-class classification.
- Parameters:
- estimatorestimator object, default=None
The base estimator to fit on self-paced under-sampled subsets of the dataset. Support for sample weighting is NOT required, but need proper
classes_andn_classes_attributes. IfNone, then the base estimator isDecisionTreeClassifier().- n_estimatorsint, default=50
The number of base estimators in the ensemble.
- k_binsint, default=5
The number of hardness bins that were used to approximate hardness distribution. It is recommended to set it to 5. One can try a larger value when the smallest class in the data set has a sufficient number (say, > 1000) of samples.
- soft_resample_flagbool, default=False
Whether to use weighted sampling to perform soft self-paced under-sampling, rather than explicitly cut samples into
k-bins and perform hard sampling.- replacementbool, default=True
Whether samples are drawn with replacement. If
Falseandsoft_resample_flag = False, may raise an error when a bin has insufficient number of data samples for resampling.- estimator_paramslist of str, default=tuple()
The list of attributes to use as parameters when instantiating a new base estimator. If none are given, default parameters are used.
- n_jobsint, default=None
The number of jobs to run in parallel for
predict().Nonemeans 1 unless in ajoblib.parallel_backendcontext.-1means using all processors. See Glossary for more details.- random_stateint, RandomState instance or None, default=None
Control the randomization of the algorithm. Within each iteration, a different seed is generated for each sampler. If the base estimator accepts a random_state attribute, a different seed is generated for each instance in the ensemble. Pass an
intfor reproducible output across multiple function calls.If
int,random_stateis the seed used by the random number generator;If
RandomStateinstance, random_state is the random number generator;If
None, the random number generator is theRandomStateinstance used bynp.random.
- verboseint, default=0
Controls the verbosity when predicting.
- Attributes:
- estimatorestimator
The base estimator from which the ensemble is grown.
- sampler_SelfPacedUnderSampler
The base sampler.
- estimators_list of estimator
The collection of fitted base estimators.
- samplers_list of SelfPacedUnderSampler
The collection of fitted samplers.
- classes_ndarray of shape (n_classes,)
The classes labels.
- n_classes_int
The number of classes.
feature_importances_ndarray of shape (n_features,)The impurity-based feature importances.
- estimators_n_training_samples_list of ints
The number of training samples for each fitted base estimators.
See also
BalanceCascadeClassifierEnsemble with cascade dynamic under-sampling.
EasyEnsembleClassifierBag of balanced boosted learners.
RUSBoostClassifierRandom under-sampling integrated in AdaBoost.
References
[1]Liu, Z., Cao, W., Gao, Z., Bian, J., Chen, H., Chang, Y., & Liu, T. Y. “Self-paced ensemble for highly imbalanced massive data classification.” 2020 IEEE 36th International Conference on Data Engineering (ICDE). IEEE, 2010: 841-852.
Examples
>>> from imbens.ensemble import SelfPacedEnsembleClassifier >>> from sklearn.datasets import make_classification >>> >>> X, y = make_classification(n_samples=1000, n_classes=3, ... n_informative=4, weights=[0.2, 0.3, 0.5], ... random_state=0) >>> clf = SelfPacedEnsembleClassifier(random_state=0) >>> clf.fit(X, y) SelfPacedEnsembleClassifier(...) >>> clf.predict(X) array([...])
Methods
Average of the decision functions of the base classifiers.
fit(X, y, *[, sample_weight])Build a SPE classifier from the training set (X, y).
get_params([deep])Get parameters for this estimator.
predict(X)Predict class for X.
Predict class probabilities for X.
score(X, y[, sample_weight])Return the mean accuracy on the given test data and labels.
set_params(**params)Set the parameters of this estimator.
- property base_estimator_
Estimator used to grow the ensemble.
- decision_function(X)
Average of the decision functions of the base classifiers.
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrices are accepted only if they are supported by the base estimator.
- Returns:
- scorendarray of shape (n_samples, k)
The decision function of the input samples. The columns correspond to the classes in sorted order, as they appear in the attribute
classes_. Regression and binary classification are special cases withk == 1, otherwisek==n_classes.
- property estimator_
Estimator used to grow the ensemble.
- property feature_importances_
The impurity-based feature importances. The higher, the more important the feature. The importance of a feature is computed as the (normalized) total reduction of the criterion brought by that feature. It is also known as the Gini importance. Warning: impurity-based feature importances can be misleading for high cardinality features (many unique values). See
sklearn.inspection.permutation_importance()as an alternative.- Returns:
- feature_importances_ndarray of shape (n_features,)
The feature importances.
- fit(X, y, *, sample_weight=None, **kwargs)
Build a SPE classifier from the training set (X, y).
- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrix can be CSC, CSR, COO, DOK, or LIL. DOK and LIL are converted to CSR.
- yarray-like of shape (n_samples,)
The target values (class labels).
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights. If None, the sample weights are initialized to
1 / n_samples.- target_labelint, default=None
Specify the class targeted by the under-sampling. All other classes that have more samples than the target class will be considered as majority classes. They will be under-sampled until the number of samples is equalized. The remaining minority classes (if any) will stay unchanged.
- n_target_samplesint or dict, default=None
Specify the desired number of samples (of each class) after the under-sampling.
If
int, all classes that have more than then_target_samplessamples will be under-sampled until the number of samples is equalized.If
dict, the keys correspond to the targeted classes. The values correspond to the desired number of samples for each targeted class.
- balancing_schedulestr, or callable, default=’uniform’
Scheduler that controls how to sample the data set during the ensemble training process.
If
str, using the predefined balancing schedule. Possible choices are:'uniform': resample to target distribution for all base estimators;'progressive': The resample class distributions are progressive interpolation between the original and the target class distribution. Example: For a class \(c\), say the number of samples is \(N_{c}\) and the target number of samples is \(N'_{c}\). Suppose that we are training the \(t\)-th base estimator of a \(T\)-estimator ensemble, then we expect to get \((1-\frac{t}{T}) \cdot N_{c} + \frac{t}{T} \cdot N'_{c}\) samples after resampling;
If callable, function takes 4 positional arguments with order (
'origin_distr':dict,'target_distr':dict,'i_estimator':int,'total_estimator':int), and returns a'result_distr':dict. For all parameters of typedict, the keys of typeintcorrespond to the targeted classes, and the values of typestrcorrespond to the (desired) number of samples for each class.
- eval_datasetsdict, default=None
Dataset(s) used for evaluation during the ensemble training process. The keys should be strings corresponding to evaluation datasets’ names. The values should be tuples corresponding to the input samples and target values.
Example:
eval_datasets = {'valid' : (X_valid, y_valid)}- eval_metricsdict, default=None
Metric(s) used for evaluation during the ensemble training process.
If
None, use 3 default metrics:'acc':sklearn.metrics.accuracy_score()'balanced_acc':sklearn.metrics.balanced_accuracy_score()'weighted_f1':sklearn.metrics.f1_score(average='weighted')
If
dict, the keys should be strings corresponding to evaluation metrics’ names. The values should be tuples corresponding to the metric function (callable) and additional kwargs (dict).The metric function should at least take 2 named/keyword arguments,
y_trueand one of [y_pred,y_score], and returns a float as the evaluation score. Keyword arguments:y_true, 1d-array of shape (n_samples,), true labels or binary label indicators corresponds to ground truth (correct) labels.When using
y_pred, input will be 1d-array of shape (n_samples,) corresponds to predicted labels, as returned by a classifier.When using
y_score, input will be 2d-array of shape (n_samples, n_classes,) corresponds to probability estimates provided by the predict_proba method. In addition, the order of the class scores must correspond to the order oflabels, if provided in the metric function, or else to the numerical or lexicographical order of the labels iny_true.
The metric additional kwargs should be a dictionary that specifies the additional arguments that need to be passed into the metric function.
Example:
{'weighted_f1': (sklearn.metrics.f1_score, {'average': 'weighted'})}- train_verbosebool, int or dict, default=False
Controls the verbosity during ensemble training/fitting.
If
bool:Falsemeans disable training verbose.Truemeans print training information to sys.stdout use default setting:'granularity':int(n_estimators/10)'print_distribution':True'print_metrics':True
If
int, print information pertrain_verboserounds.If
dict, control the detailed training verbose settings. They are:'granularity': corresponding value should beint, the training information will be printed pergranularityrounds.'print_distribution': corresponding value should bebool, whether to print the data class distribution after resampling. Will be ignored if the ensemble training does not perform resampling.'print_metrics': corresponding value should bebool, whether to print the latest performance score. The performance will be evaluated on the training data and all given evaluation datasets with the specified metrics.
Warning
Setting a small
'granularity'value with'print_metrics'enabled can be costly when the training/evaluation data is large or the metric scores are hard to compute. Normally, one can set'granularity'ton_estimators/10(this is used by default).
- Returns:
- selfobject
- get_params(deep=True)
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- predict(X)
Predict class for X.
The predicted class of an input sample is computed as the class with the highest mean predicted probability. If base estimators do not implement a
predict_probamethod, then it resorts to voting.- Parameters:
- X{array-like, sparse matrix} of shape (n_samples, n_features)
The training input samples. Sparse matrices are accepted only if they are supported by the base estimator.
- Returns:
- yndarray of shape (n_samples,)
The predicted classes.
- predict_proba(X)
Predict class probabilities for X.
The predicted class probabilities of an input sample is computed as the mean predicted class probabilities of the base estimators in the ensemble. If base estimators do not implement a
predict_probamethod, then it resorts to voting and the predicted class probabilities of an input sample represents the proportion of estimators predicting each class.- Parameters:
- X{array-like, sparse matrix} of shape = [n_samples, n_features]
The training input samples. Sparse matrices are accepted only if they are supported by the base estimator.
- Returns:
- parray of shape = [n_samples, n_classes]
The class probabilities of the input samples.
- score(X, y, sample_weight=None)
Return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
- Parameters:
- Xarray-like of shape (n_samples, n_features)
Test samples.
- yarray-like of shape (n_samples,) or (n_samples, n_outputs)
True labels for X.
- sample_weightarray-like of shape (n_samples,), default=None
Sample weights.
- Returns:
- scorefloat
Mean accuracy of
self.predict(X)wrt. y.
- set_params(**params)
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Examples using imbens.ensemble.SelfPacedEnsembleClassifier
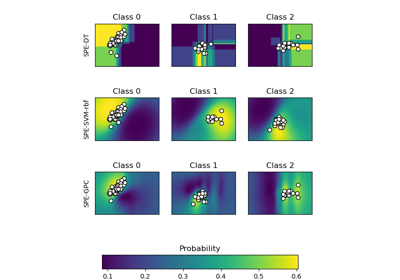
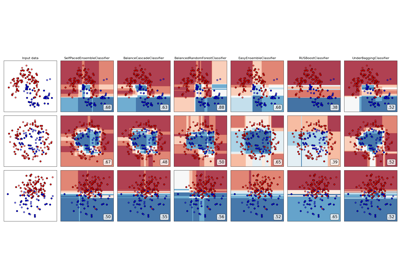
Plot probabilities with different base classifiers